-
开放科学(资源服务)标识码(OSID):

-
20世纪初,弗洛尔提出的基因对基因假说作为极具开创性的重要概念之一,为植物免疫领域众多深入研究开辟了道路[1]。多年来,这一假说指导着现代植物免疫研究。在过去的30多年里,我们见证了分子生物学领域的快速发展,特别是在分子植物—微生物互作领域。随着对编码免疫受体、病原体相关分子模式(Pathogen-Associated Molecular Pattern,PAMP)、损伤相关分子模式(Damage-Associated Molecular Patterns,DAMP)、病原体无毒(Avriulence,Avr)基因、植物抗性(Resistance,R)基因以及植物免疫反应关键信号成分等的鉴定与表征,植物免疫学不断发展,进一步提升了我们对植物与微生物互作的认识与理解。
植物不断受到各种生物(包括病毒、细菌、真菌、卵菌等)的侵染挑战。在与病原体长期竞争的过程中,植物进化出先天免疫系统以抵抗病原体的攻击。植物的先天免疫反应由位于细胞表面的模式识别受体(Pattern Recognition Receptors,PRRs)和细胞内NB-LRR受体(Nucleotide-Binding Domain Leucine-Rich Repeat Containing Receptors,NLRs)启动,从而分别介导PTI免疫(PAMP-Triggered Immunity)和ETI免疫(Effector-Triggered Immunity)[2-3]。PTI是植物初级防御反应之一,在PRRs识别PAMP或DAMP后被激活[4];而更深一层的先天免疫由NLRs介导的ETI防御反应触发[5]。ETI通常更为强烈,并常伴随被称为过敏反应(Hypersensitive Response,HR)或过敏性细胞坏死反应的程序性细胞死亡,从而有效增强寄主抗性。本综述以NLRs介导的植物免疫为出发点,概述植物免疫体系中NLRs的研究进展与激活机制,以及其激活后的免疫信号传导机制,旨在更好地理解不同免疫受体如何识别具有不同攻击策略的病原体,为进一步开发新型作物抗病育种策略提供理论依据。
HTML
-
NLRs属于具有多个结构域的信号转导ATPase(Signal Transduction ATPases with Numerous Domains,STANDs)大家族[6]。与大多数STAND蛋白一样,植物NLRs是一类分子开关,其在静止状态下与ADP结合而处于非活性构象,在感知到非自身或经修饰的自身信号后可被条件性激活并启动免疫信号传导[7]。植物中大多数NLRs包含3个结构域,即N端可变结构域、中间核苷酸结合与寡聚化结构域(Nucleotide-Binding and Oligomerization Domain,NOD或NB)以及C端富含亮氨酸重复序列结构域(Leucine-Rich Repeat,LRR)[8-9]。此外,一些NLRs还整合了额外的非规范结构域,称为NLR整合结构域(NLR-integrated Domains,NLR-IDs)。N端结构域通常被认为是信号结构域,在NLRs受激活后介导下游的程序性细胞死亡反应[10]。中间NOD结构域也称NB-ARC结构域(Nucleotide-Binding Adaptor Shared by APAF-1,Certain R Gene Products,and CED-4),其在NLRs由非活性状态向活性状态转换的过程中发生决定性的构象变化[11-12]。C端LRR结构域决定对病原体的识别特异性;此外,该结构域还介导分子内的自抑制相互作用,当自抑制状态被解除后,N端结构域随即介导下游免疫信号传导[7, 13]。
植物NLRs可依据其N端结构域分为不同类型,这些类型与NB-ARC结构域的系统发育关系相一致,提示其具有深远的进化起源[14]。在被子植物中已发现4类主要的N端信号结构域特征,卷曲螺旋(Coiled-Coil,CC)型、抗白粉病8样结构域(Resistance to Powdery Mildew 8-like Domain,RPW8)型(亦称CCR型)、G10-type CC型(亦称CCG10型)以及Toll/白细胞介素-1受体/抗性蛋白(Toll/interleukin-1 Receptor/Resistance Protein,TIR)型;而在不开花植物中,NLRs还可携带其他类型的N端结构域,如α/β水解酶结构域和激酶结构域[7]。部分CC结构域参与效应子识别,可直接与效应子(如Avr-Pik)及病原体效应子靶标的“守卫”蛋白(Guardee Protein,如RIN4)相互作用;CC和RPW8结构域中的α螺旋已被证实可形成防御信号传导所需的阳离子通道[15-16];TIR结构域可自我关联,或与配对TNLs(TIR型NLRs)中的TIR结构域相互关联以完成自身激活;TIR结构域在寡聚化后表现出NAD酶活性,生成v-cADPR(variant-cyclic ADP-ribose),且该酶活性为诱导过敏性细胞死亡所必需[17-18];此外,TIR结构域还具有2′,3′-cAMP/cGMP合成酶活性,该催化活性同样为诱导过敏性细胞死亡所必需[19]。
通常情况下,成簇的NLR变体存在于多态性位点中,这些位点通过串联重复、不等交换事件以及插入和突变等过程产生[20-22]。进化基因组学研究表明,植物免疫受体基因(NLRs及部分PRRs)是植物基因组中进化速度最快的基因之一[23-24]。不同NLRs的LRR结构域往往呈现最大的变异性,这与其在识别快速演化的病原体效应子方面的功能相一致[25]。
NLR编码基因广泛存在于陆地植物基因组中,但不同植物物种间NLR编码基因数量差异显著。例如,拟南芥的NLR编码基因超过160个,而小麦可超过1 500个。此外,即使在同一物种内,NLR编码基因在数量和序列层面也可表现出显著多样性[8, 22-23]。CNLs(CC型NLRs)、TNLs和RNLs(RPW8型NLRs)在基部被子植物(如无油樟属Amborella和睡莲属Nymphaea)中均存在,而在大多数单子叶植物基因组中缺乏TNLs,提示该类基因的丢失可能发生在单子叶植物与双子叶植物分化之前[26-28]。TNLs的缺失还伴随其信号成分Enhanced Disease Susceptibility 1(EDS1)、Phytoalexin Deficient 4(PAD4)和Senescence-Associated Gene 101(SAG101)的丢失,这些信号成分的缺失可能驱动了一些被子植物谱系中TNLs的收缩[27-28]。此外,NLR基因家族还经历了谱系特异性的共同扩张或共同收缩,其原因尚不明确,但有观点认为这可能与病原体压力及生态特化相关[24, 27-28]。
-
植物免疫系统中,PRRs介导的免疫与NLRs介导的免疫并非相互独立,而是相互协同并彼此增强[3, 29-30]。有研究报道,NLRs的激活可导致多个PRR信号组分在转录本和蛋白质水平上的积累,从而增强并延长PRRs介导的免疫反应[31]。值得注意的是,NLRs的激活同样依赖PRRs介导的免疫。以丁香假单胞菌(Pseudomonas syringae)分泌的效应子AvrRpt2为例,该效应子通过激活NLR蛋白RPS2触发寄主ETI反应,但在PRR或PRR共受体缺陷突变体(如fls2、pepr3、fls efr cerk1以及bak1-5 bkk1 cerk1等基因突变体)中该过程失效[29-30]。此外,PRRs的激活会促进多个NLRs及含有EP结构域蛋白的转录本积累,且PRRs介导的免疫也在一定程度上依赖EP蛋白与辅助NLRs[32]。因此,PRRs与NLRs之间的互作对赋予植物有效抗病能力至关重要,但其协同增强的分子机制仍有待进一步阐明。
-
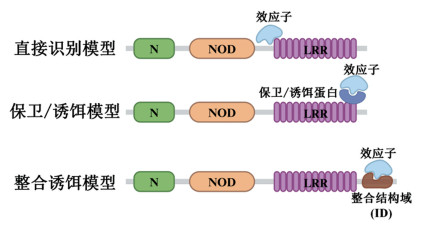
近年来,关于NLRs作为病原体激活的分子开关以对抗植物病害的研究取得了显著进展。NLRs激活的第一步是对病原体效应子的特异性识别。根据当前研究,该识别过程主要包括直接识别模型、保卫模型、诱饵模型以及整合诱饵模型4种分子模型(图 1)[33-34]。
直接识别模型是指NLRs通过其LRR结构域直接与病原体效应子结合。例如,拟南芥TNL蛋白RPP1(Recognition of Peronospora parasitica 1)和本氏烟TNL蛋白Roq1(Recognition of XopQ 1)可直接与其同源效应子配体结合[35-37]。类似地,小麦CNL蛋白Sr35也可直接结合其同源效应子AvrSr35[38-39]。
保卫模型与诱饵模型在分子作用原理上相近,二者均通过NLRs监控特定宿主蛋白(或蛋白质结构域)的功能或物理完整性来激活免疫应答,即NLRs扮演“守卫”角色:当检测到受保护蛋白(结构域)被病原体效应子修饰或破坏时,迅速启动级联免疫信号传导[40]。两种模型的核心差异在于被守卫对象的生物学特性。保卫模型中,NLRs守卫的是参与宿主防御的蛋白;诱饵模型中,NLRs守卫的是效应子靶向的诱饵蛋白,这类诱饵蛋白本身不具备抗病活性,而是在结构上模仿具有抗性功能的宿主蛋白以引发病原体攻击,因此诱饵蛋白可作为毒力相关活动的“传感器”[41]。例如,拟南芥RPS2与RPM1均监测RIN4蛋白的状态;RIN4是多种不相关细菌III型效应子的靶标。RPS2在效应子AvrRpt2的蛋白酶活性切割RIN4时被激活;RPM1则在检测到效应子AvrB或AvrRpm1介导的RIN4翻译后修饰扰乱其磷酸化转换状态时被激活[42-44]。RPS5监测类受体胞质激酶(Receptor-Like Cytoplasmic Kinase,RLCK)PBS1的状态,当PBS1被AvrPphB蛋白酶切割时,RPS5被激活;在此过程中,PBS1发挥诱饵作用[45]。此外,CNL蛋白ZAR1(HopZ-Activated Resistance 1)也是典型代表。ZAR1是少数在远缘植物物种中仍存在同源体的NLRs之一,它通过中间假激酶(如ZED1或RKS1)监测多种类RLCK蛋白的状态,从而被激活并触发免疫反应[46-47]。
整合诱饵模型是一种额外的识别模式,其提出源于对非典型NLRs蛋白结构与功能的解析。这类NLRs携带效应子感知所需的非规范整合域(NLR-IDs),并利用IDs作为诱饵以直接或间接识别效应子。例如,NLR-ID的典型代表RRS1(Resistance to Ralstonia Solanacearum 1)包含C端WRKY结构域作为ID,并能与另一NLR蛋白RPS4(Resistance to Pseudomonas Syringae 4)形成受体复合物(RPS4/RRS1)。RRS1的ID可与细菌效应子PopP2(青枯雷尔氏菌分泌的乙酰转移酶效应子)或AvrRps4(丁香假单胞菌分泌的效应子)直接结合,从而激活RPS4/RRS1介导的免疫反应[48-49]。
-
结构生物学与冷冻电子显微镜(cryo-EM)研究揭示了NLR蛋白激活后的构象变化。在多数情况下,NLRs的NOD结构域在非活性状态下结合ADP;当识别到病原体效应子后,蛋白构象发生改变,使ADP与ATP交换;ATP结合状态下,NOD结构域通过寡聚化形成聚合体,促使N端信号结构域聚集并启动信号传导[50]。NLRs激活后形成的聚合体被称为抗病小体(Resistosomes)[51]。
植物免疫系统中首个被解析的抗病小体结构来自CNL蛋白ZAR1。该蛋白通过守卫PTI所需的RLCK或其诱饵来识别细菌效应子。野油菜黄单胞菌(Xanthomonas campestris)产生的AvrAC效应子可使BIK1、PBL2等RLCK发生尿苷酰化,即向RLCK添加尿苷单磷酸基团(UMP),进而抑制RLCK活性并损害植物PTI免疫;ZAR1与假激酶RKS1形成异源二聚体复合物(RKS1-ZAR1),监测由AvrAC引起的诱饵PBL2尿苷酰化,并与修饰后的PBL2(PBL2^UMP)相互作用形成ZAR1-RKS1-PBL2UMP复合物,触发ADP/ATP交换并驱动五聚体形成。该五聚体以漏斗状结构在质膜上形成阳离子通道,促进Ca2+内流,进而激活ETI相关基因表达并启动程序性细胞死亡[12, 15]。此外,小麦CNL蛋白Sr35在直接识别病原效应子AvrSr35后也可形成五聚体结构,其N端与ZAR1五聚体表现出相似的寡聚化特征。这些发现表明,CNL抗病小体及其离子通道活性具有一定的进化保守性,并揭示了抗病小体在触发细胞死亡与抗病反应中的关键作用[39]。
拟南芥TNL蛋白RPP1识别病原体特定ATR1变体并与之直接结合,随后RPP1寡聚形成四聚体抗病小体。该构象变化使N端TIR结构域聚集,从而激活其NAD酶活性,将NAD+水解为可被EDS1-PAD4或EDS1-SAG101异源二聚体感知的小信号分子[35]。近年来,关于植物NLRs激活前后构象变化的研究进展,显著加深了我们对NLRs识别与激活机制的理解。
-
目前,“NLR受体网络”概念被广泛用于阐述多个NLRs协同触发免疫信号的过程。在该网络中,负责识别与检测病原体效应子的NLRs被称为传感器NLRs(sensor NLRs,sNLRs),而将sNLRs的激活信号向下游转导的NLRs被称为辅助NLRs(helper NLRs,hNLRs)[52]。迄今已鉴定出NRG1(N Required Gene 1)、ADR1(Activated Disease Resistance 1)和NRC(NLR Required for HR-Associated Cell Death)3类hNLRs。其中,NRG1与ADR1的N端均含RPW8结构域,亦称RNLs;NRG1是最早被确认的hNLR成员之一[52-55]。NRG1类hNLRs主要在TNLs下游发挥作用,而ADR1类hNLRs则参与TNL与CNL介导的免疫[56-57]。
NRG1与ADR1需要通过与EDS1形成复合物以实现免疫的完全激活。EDS1在多种TNL介导的免疫反应中充当中央信号枢纽。在拟南芥及其他双子叶植物中,EDS1-PAD4复合物与ADR1类hNLRs协同以激活转录依赖性防御,而EDS1-SAG101复合物与NRG1类hNLRs协同以促进防御过程中细胞死亡[58]。EDS1-SAG101-NRG1与EDS1-PAD4-ADR1两条信号模块已被鉴定为TNLs下游免疫信号通路的关键组成。当TNL被病原效应子激活并形成四聚体后,其TIR结构域的NAD酶活性产生小核苷酸衍生信号分子(pRib-AMP/ADP或ADPr-ATP/di-ADPR),分别作用于EDS1-PAD4或EDS1-SAG101异源二聚体,促进其与ADR1或NRG1结合,进而形成ADR1与NRG1抗病小体(即辅助RNLs抗病小体)。该复合体可在质膜上充当Ca2+通道,引发后续一系列免疫信号[50, 52]。
另一类hNLR-NRC在茄科植物中支持近一半sNLRs的功能[59]。NRC家族包括NRC1、NRC2、NRC3、NRC4及一个非典型成员NRCX。多个NRC成员之间存在功能冗余,部分sNLRs介导的免疫需要NRC2、NRC3与NRC4共同参与,而另一些则仅需NRC2与NRC3,或仅需NRC4[50]。NRC蛋白N端具有一个保守且功能等效的N端基序(MADA基序),该基序由受体一端约21个氨基酸组成,并在约20%的CNLs中保守。值得注意的是,MADA基序与ZAR1蛋白N端α1螺旋约有50%的序列相似性,而后者在ZAR1抗病小体激活过程中会发生关键构象转换[60]。sNLRs检测到配体后激活NRC,促使形成NRC寡聚体,而sNLRs被排除在该寡聚体之外。最新研究表明,NRC4在被sNLR蛋白Bs2及致病效应子AvrBs2激活后可组装为六聚体抗病小体,该构象变化同样通过促进Ca2+进入细胞质以触发免疫反应[61-62]。
细胞内Ca2+内流长期以来被认为是防御反应与NLRs激活的重要标志之一[50, 63]。已有研究证实,激活的NLRs可作为Ca2+通道发挥作用,如直接激活的CNLs(ZAR1、Sr35)以及由TNLs激活后产生的小信号分子间接激活的RNLs(NRG1/ADR1)均可引发Ca2+通道活性,从而通过钙信号调控植物免疫反应。
3.1. NLR蛋白对病原体效应子的识别
3.2. 识别激活后NLRs的结构变化
3.3. NLR受体网络的免疫信号传导机制
-
作物病害是造成粮食产量大幅下降的主要因素之一,影响全球粮食安全。选育抗病品种是农业病害防治中最经济、最环保的措施。深入理解植物—病原体互作及免疫机制,对于制定提升抗病性的作物育种策略至关重要。NLRs介导的植物免疫机制研究为阐明植物与病原体互作提供了重要见解,也为作物抗病育种提供了新的思路与策略。目前,具有已知效应子的NLRs数量已超过140个,且新型NLRs仍在不断被鉴定[5]。NLRs激活机制(尤其是抗病小体)的发现,使植物防御网络图景更加清晰,并为工程化抗病育种提供了新的线索。
植物免疫受体的生物工程改造有望成为获得新型抗病性状的关键策略,以应对植物病原体对全球粮食安全日益加剧的威胁。在该领域中,NLR受体的结构域工程展现出巨大潜力。以稻瘟病防控为例,Zdrzalek等[64]将水稻NLR蛋白Pikm-1(通常通过识别效应子AVR-PikD抵抗稻瘟病)中的整合HMA结构域替换为来自水稻蛋白OsHIPP43(该蛋白为稻瘟病菌效应蛋白Pwl2的靶标)的HMA结构域,构建的新型嵌合受体使水稻对携带PWL效应蛋白的稻瘟病菌分离株产生广谱抗性。这一创新性改造为NLR蛋白的定制化设计提供了重要支撑,并为定向扩展病原识别谱提供了可拓展平台。未来,在进一步系统解析NLR蛋白功能及进化规律的基础上,可整合基因编辑与合成生物学等技术,精准设计具有多靶标识别能力的NLR受体阵列,从而培育兼具广谱性与持久性的抗病作物品种。此外,将NLRs介导的特异性免疫调控与RNA干扰、微生物组调控等新型防病策略相结合,有望构建多层次植物防御网络。这种协同增效模式不仅可延缓病原体的适应性进化,也有助于减少对化学农药的依赖,为建立可持续农业体系提供创新解决方案。




 DownLoad:
DownLoad: