-
开放科学(资源服务)标识码(OSID):

-
中华按蚊Anopheles sinensis隶属节肢动物门(Arthropod)、昆虫纲(Insecta)、双翅目(diptera)、蚊科(Culicidae)、按蚊属Anopheles,是疟疾、马来丝虫病、乙型脑炎等疾病的重要媒介蚊虫,在我国和东南亚国家广泛分布[1-3]。目前,在媒介蚊虫防控中化学防治是主要手段。由于拟除虫菊酯杀虫剂具有高效、低毒等优势,成为了防控疟疾媒介蚊虫最常用的杀虫剂[4-6]。但随着拟除虫菊酯杀虫剂被长期广泛使用,学者们发现其药效明显降低,蚊虫对该类药剂产生了不同程度的抗性[7-10]。
细胞色素P450酶是参与代谢外源性、内源性化合物以及各种杀虫剂的重要解毒酶系[11],可以与拟除虫菊酯杀虫剂分子基团发生氧化、脱氨、羟基化等化学反应,从而使拟除虫菊酯杀虫剂失去毒性,因此细胞色素P450酶在拟除虫菊酯杀虫剂抗性机制中起着重要作用[12]。先前研究发现,蚊虫CYP6、CYP9、CYP4亚家族成员与杀虫剂抗性有关,其中CYP6亚家族成员被认为与抗性密切相关[13]。例如:冈比亚按蚊的CYP6P3和CYP6M2基因与拟除虫菊酯类杀虫剂抗性密切相关[14];淡色库蚊的CYP6AA9基因在溴氰菊酯抗性品系中上调表达,其被RNAi(RNA干扰)沉默之后导致淡色库蚊的死亡率增加[15];埃及伊蚊的CYP6BB2、CYP6M11、CYP9J23基因在拟除虫菊酯类杀虫剂抗性中上调表达[16]。Yan等[17]通过RNA-seq和RT-qPCR验证发现,CYP6Z2、CYP6P3v1、CYP6P3v2、CYP9J5、CYP306A1基因在拟除虫菊酯抗性中显著上调表达,可能与拟除虫菊酯抗性相关。在实验室前期研究中,Guo等[18]发现3个(CYP9K1、CYP6P3v1、CYP9J10) P450基因被RNAi沉默之后,降低了中华按蚊对拟除虫菊酯类杀虫剂的抗性,证实了这3个P450基因与拟除虫菊酯抗性相关。研究表明,P450基因上调表达可能源于其调控区DNA序列发生核苷酸插入、缺失或者碱基替换等变异,这些变异可能会增强调控元件和转录因子的结合能力,或者促使新的调控元件产生,或者破坏沉默子的功能性结合位点[19-20]。解毒酶基因编码区的变异可以修饰蛋白酶结构,增强解毒酶活性,从而提高其对杀虫剂的代谢解毒效率[21-23]。目前,昆虫P450基因编码区变异对杀虫剂产生抗性的报道较少,之前在牧草盲蝽Lygus pratensis抗苄氯菊酯品系中,发现CYP6X1基因编码区存在大量的核苷酸变异,其变异与抗性的关联性需进一步研究[24]。在褐飞虱的吡虫啉抗性品系中,发现CYP6ER1编码区存在突变,这可能是褐飞虱对杀虫剂产生抗性的重要因素[25]。
SNP是物种适应环境变化和进化的重要分子基础,特定的SNP可能影响基因表达,从而影响生物对环境的适应能力。目前,全基因组水平检测蚊虫抗性相关的单核苷酸多态性研究较少[26]。
本研究对36个中华按蚊个体进行遗传变异检测,并对P450基因的SNPs数据进行进化树分析、主成分分析、遗传结构分析、遗传变异分析。其中,重点分析了中华按蚊拟除虫菊酯抗性相关P450基因的遗传变异特征,为深入解析中华按蚊拟除虫菊酯抗性相关P450基因表达调控和解毒酶活性机制奠定了数据基础。
HTML
-
在重庆、云南、安徽3个省(市)的稻田中采集4龄按蚊幼虫[27],并在当地养蚊室内饲养至成虫。室内饲养条件为27 ℃、相对湿度80%。随后,提取蚊虫足部进行物种的分子鉴定[28]。雌性成蚊羽化72 h后,采用世界卫生组织(WHO)推荐的药膜生物测定法进行抗性检测,将雌性成蚊暴露在含有0.05%溴氰菊酯药膜接触桶中60 min,随后将其转移到恢复桶中,若蚊虫在60 min内死亡或者机械刺激无反应,则判定为拟除虫菊酯敏感个体;24 h后统计蚊虫的死亡率,存活个体判定为拟除虫菊酯抗性个体。敏感品系成蚊的相对敏感基线LC50(半致死浓度)为0.006 7 mg/L,云南抗性品系(YN-FR)、重庆抗性品系(CQ-FR)、安徽抗性品系(AH-FR)的抗性倍数分别为500倍、800倍、800倍,均达到国家标准的高抗水平[29]。最终,从重庆、云南、安徽3个省(市)分别收集6只拟除虫菊酯抗性成蚊(Field Pyrethroid-Resistant Strain,FR)和6只拟除虫菊酯敏感成蚊(Field Pyrethroid-Susceptible Strain,FS),共36个样本用于后续分析。
-
将36个中华按蚊野外样本送往华大公司进行测序,并采用Illumina HiSeq2000平台测序获得重测序数据,中华按蚊的参考基因组(GCA-000441895.2)由重庆师范大学昆虫与分子生物学研究所提供[30-31]。
-
使用fastp软件[32]对原始测序数据(Raw reads)进行质量控制与过滤,获得高质量测序读段(Clean reads)。然后,使用BWA软件[33]将36个中华按蚊样本的高质量测序读段与参考基因组进行比对。通过SAMtools(处理高通量测序比对数据的软件集)[34]将比对生成的SAM(存储测序序列与参考基因组比对结果的文本格式)文件转换为排序后的BAM(SAM的二进制压缩格式)文件,并去除PCR(聚合酶链式反应)重复序列,最终获得用于后续分析的比对结果文件。
-
使用GATK[35]对36个中华按蚊样本的比对结果进行SNPs和InDels变异检测。为确保数据质量,对原始变异位点进行过滤,并使用ANNOVAR软件[36]进行注释分析。
-
前期基于中华按蚊基因组注释蛋白库,使用BLASTP和Hmmsearch两种方法,获得中华按蚊P450基因[17]。再使用VCFtools软件[37]对GATK过滤后的变异结果进一步筛选,剔除次要等位基因频率(maf)<0.05的位点、去除基因组型缺失率超过20%的位点,同时仅保留二等位SNPs变异位点。基于中华按蚊基因组注释文件,使用Python脚本提取P450基因家族成员的注释信息。最后,根据P450基因在基因组中的起始和终止位置,使用VCFtools(参数:-bed)提取All.filtered.vcf文件中的变异位点[38],作为P450基因家族成员的SNPs数据,用于后续群体结构分析。
-
使用PHYLIP软件中的邻近法(Neighbor-Joining Methods)构建系统发育树(F84模型,100次Bootstrap重复),并使用Ggtree进行可视化展示。
-
基于36个中华按蚊样本的遗传变异数据,使用GCTA软件[39]进行主成分分析,并根据主成分聚类结果将36个中华按蚊样本划分为不同的遗传亚群。
-
使用Admixture软件[40]进行群体遗传结构分析。通过最大似然估计法确定最优群体数目K(预设范围2-N),并使用贝叶斯算法计算每个K值下的群体分群。
-
36个样本分为6个组,即安徽抗性组AH-FR、安徽敏感组AH-FS、重庆抗性组CQ-FR、重庆敏感组CQ-FS、云南抗性组YN-FR、云南敏感组YN-FS。每组6个样本,且相同地区的抗性组和敏感组比较,如安徽抗性组AH-FR与安徽敏感组AH-FS比较,另外两个地区也是如此。将组内基因型一致,但组间基因型存在差异的位点定为差异位点。筛选差异位点的标准为:在安徽抗性组(AH-FR)中,6个样本中有80%的样本(即5个样本)具有一致的基因型(记为该组的优势基因型A);同时在安徽敏感组(AH-FS)中,6个样本中也要有80%的样本(即5个样本)具有一致的基因型(记为该组的优势基因型B)。仅当A与B不同时,该位点才被定义为差异位点。然后,对这些位点进行注释,确定其所在基因位置及突变类型。
1.1. 蚊虫来源
1.2. 数据来源
1.3. 低质量数据过滤和序列比对
1.4. 群体SNPs和InDels变异检测及注释
1.5. 群体结构分析的SNPs
1.6. 进化树分析
1.7. 主成分分析
1.8. 群体遗传结构分析
1.9. 筛选中华按蚊拟除虫菊酯抗性相关P450基因的差异SNPs
-
序列比对结果如表 1所示。由表 1可知,36个样本的比对率为78.26%~98.64%,平均测序深度为8.769×~27.817×,说明样本建库良好;参考基因组的覆盖度为89.73%~97.31%,说明比对情况良好,可用于下一步群体遗传结构分析。
-
P450基因的变异数量统计结果如表 2所示。36个中华按蚊样本的P450基因家族共检测到59 070个变异位点,其中SNPs数量为46 952个,平均每kb区间内鉴定出0.68个SNPs位点;InDels数量为12 118个,平均每kb区间内鉴定出0.19个InDels位点。
P450基因SNPs和InDels的注释统计结果如表 3所示。SNPs被注释到基因间区(intergenic regions)的数量为62 011个,内含子区域数量为45 750个,下游区域SNPs数量为11 416个,上游区域SNPs数量为11 795个,外显子区域SNPs数量为13 971个,5'端非翻译区(5'UTR)SNPs数量为1 085个,3'端非翻译区(3'UTR)SNPs数量为1 478个,剪切位点区域(splicing)SNPs数量为25个。InDels被注释到基因间区(intergenic regions)的数量为9 238个,内含子区域InDels数量为7 487个,上游区域InDels数量为1 976个,3'端非翻译区(3'UTR)InDels数量为248个,外显子区域InDels数量为224个,下游区域InDels数量为1 831个,5'端非翻译区(5'UTR)InDels数量为152个,剪切位点区域(splicing)InDels数量为13个。由此可见,中华按蚊P450基因的SNPs和InDels主要分布在基因间区(intergenic regions),其次是内含子区。
-
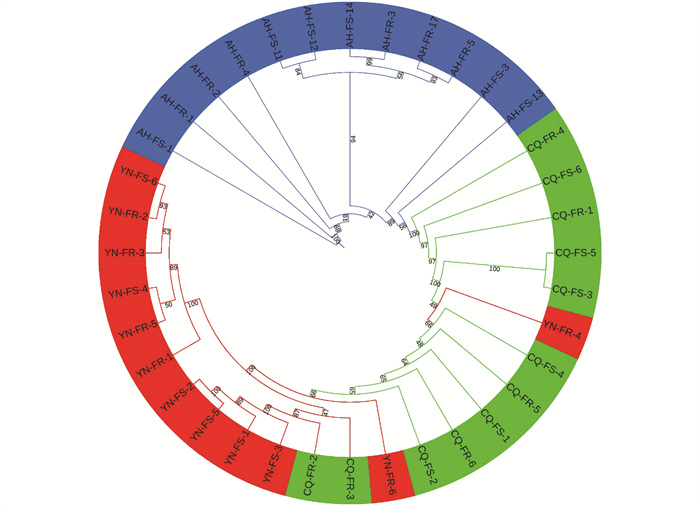
基于P450-SNP的中华按蚊系统进化树分析结果如图 1所示。36个中华按蚊样本按照地理种群聚类为3组,种群间的遗传差异和地理区域相对应。安徽12个样本中AH-FS-13、AH-FR-2自展值较低(<70%);而其他样本以较高的自展值聚类(>80%),说明这些样本可能来自共同祖先。云南12个样本中YN-FR-5、YN-FR-4、YN-FS-4、YN-FR-3自展值较低(<70%);而其他样本都以较高的自展值聚类(>80%),说明这些样本可能来自共同祖先。重庆12个样本中CQ-FR-3、CQ-FS-2、CQ-FR-6、CQ-FS-1、CQ-FR-5、CQ-FS-4自展值较低(<70%);而其他样本都以较高的自展值聚类(>80%),说明这些样本可能来自共同祖先。CQ-FS-5与CQ-FS-3以自展值100%聚类在一起,说明它们可能来自共同祖先。CQ-FR-2和CQ-FR-3与重庆种群其他样本显著分离,表明重庆种群的群内遗传差异较大,含有不同的祖先来源。
-
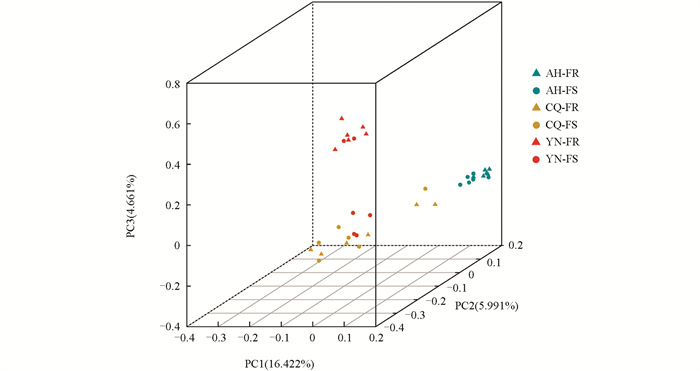
主成分分析结果如图 2所示。36个中华按蚊样本聚类与地理分布基本一致,其中云南(YN)和安徽(AH)种群各自形成独立聚类,而重庆(CQ)种群内部个体分散,表明其遗传多样性较高。此外,图 2中PC1、PC2和PC3分别代表第一、第二和第三主成分,其后方括号内的百分比(16.422%、5.991%、4.661%)为各主成分的方差贡献率,由eigenval.xls特征值文件计算得出。由此可知,主成分分析结果与系统进化树的分析结果一致。
-
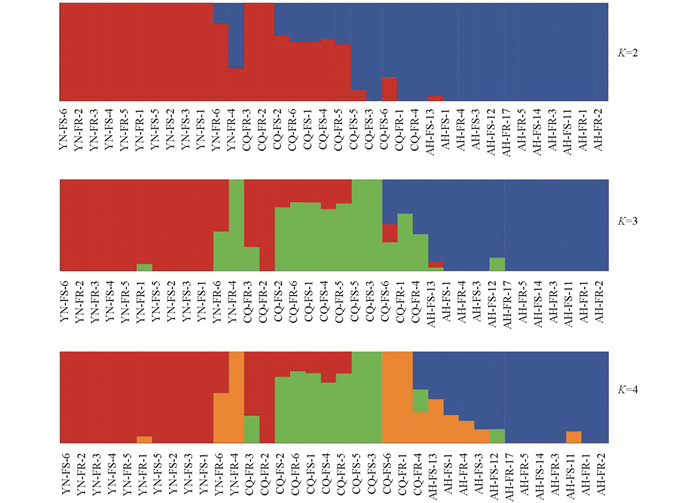
群体遗传结构分析结果如图 3所示。随着设定的祖代群体数K从2逐步提高至4,中华按蚊36个样本的群体结构呈现相应的动态变化。当K=2时,36个中华按蚊可划分为两个群体;当K=3时,重庆种群个体从云南和安徽种群中分化出来,36个中华按蚊样本可划分为3个群体;当K=4时,重庆种群的群内差异较大,产生了一个亚群。综上,群体遗传结构分析结果与系统进化树分析和主成分分析结果一致。
-
中华按蚊拟除虫菊酯抗性相关的P450基因在重庆、云南、安徽抗性种群中的差异SNPs分别如表 4、表 5、表 6所示。结合实验室前期研究发现的3个(CYP9K1、CYP6P3v1、CYP9J10) P450基因被RNAi干扰后,降低了中华按蚊对拟除虫菊酯类杀虫剂的抗性。基于此,本研究重点分析这3个基因的遗传变异。CYP9K1基因在重庆、云南、安徽抗性种群中检测到21个差异SNPs位于基因间区,1个同义替换(L493L)位于外显子区域。CYP6P3v1基因在云南抗性种群中检测到1个同义替换(L118L)和1个差异SNPs,分别位于外显子区域和内含子区域。CYP9J10基因在重庆抗性种群中检测到1个非同义替换(S162N),位于外显子区域。综上,CYP6P3v1存在外显子区域同义替换(L118L)和内含子区域SNPs;CYP9J10存在外显子区域非同义替换(S162N);而CYP9K1主要存在基因间区SNPs。此外,这些基因均属于CYP6和CYP9家族成员。
2.1. 序列比对结果分析
2.2. P450基因的SNPs和InDels变异鉴定结果与注释分析
2.3. 进化树分析结果
2.4. 主成分分析结果
2.5. 群体遗传结构分析结果
2.6. 中华按蚊拟除虫菊酯抗性相关P450基因的差异SNPs
-
本研究通过全基因组重测序分析发现,中华按蚊P450基因的SNPs和InDels主要分布于基因间区,这可能与基因间区在基因组中所占比例较高有关。通过进化树分析、主成分分析、群体遗传结构分析,发现36个中华按蚊的聚类关系与其地理分布基本一致。种群间的遗传差异与地理区域相对应,这与以往的研究结果一致,即中华按蚊的遗传距离和地理距离呈正相关,可能存在地理隔离现象[41]。
本研究首次发现,CYP6P3v1存在外显子区域同义替换(L118L)和内含子区域SNPs,CYP9J10存在外显子区域非同义替换(S162N),而CYP9K1主要存在基因间区SNPs。此外,这些基因均属于CYP6和CYP9家族成员,该结果与埃及伊蚊差异SNPs检测结果一致[26]。
Dusfour等[42]报道埃及伊蚊(Ae.aegypti) CYP9J10、CYP9J9、CYP6BB2、CYP6M11和CYP6N12基因的表达与拟除虫菊酯类杀虫剂抗性相关。Yan等[17]通过RNA-seq转录组测序和RT-qPCR验证发现,中华按蚊CYP9K1基因在重庆和安徽2个地理种群的拟除虫菊酯抗性品系中上调表达;中华按蚊CYP6P3v1基因在重庆、云南、安徽3个地理种群的拟除虫菊酯抗性品系中均呈现上调表达。已有报道催命按蚊CYP9K1与拟除虫菊酯类杀虫剂抗性相关[43]。中华按蚊的CYP6P3v1已被证实与拟除虫菊酯类杀虫剂抗性有关[18]。
以往的研究报道,P450基因的代谢抗性主要是由基因上调表达和解毒酶活性增加所致[44-45]。蚊虫对拟除虫菊酯的抗性是由解毒酶基因编码区变异产生的[46]。昆虫P450基因编码区的遗传变异可导致杀虫剂抗性,例如埃及伊蚊(Ae.aegypti)抗性品系中的CYP6、CYP9和CYP12基因家族均存在与抗性相关的突变位点[47]。非翻译区的突变能调控基因表达,进而导致杀虫剂抗性[48]。研究表明,同义替换SNP可通过影响mRNA稳定性、翻译效率或剪接过程参与基因表达调控[49]。内含子区域的突变可能改变内含子的剪切形式,产生不同的剪切体,由于这些剪切体稳定性不同,从而影响基因表达水平[23]。Amichot等[50]在黑腹果蝇(Drosophila melanogaster)的双对氯苯基三氯乙烷(DDT)中发现3个非同义替换(R335S、L336V、V476L)位于CYP6A2活性位点附近,这些突变位点与DDT的代谢抗性有关。本研究发现重庆和安徽抗性种群的CYP9K1仅存在一种变异类型(即基因间区SNPs);云南抗性种群的CYP6P3v1存在两种变异类型(即外显子区域同义替换、内含子区域SNPs);重庆抗性种群的CYP9J10存在一种变异类型(即外显子区域非同义替换)。
本研究外显子区域同义替换(L118L)、内含子区域及基因间区SNPs可能正向调控CYP6P3v1和CYP9K1的表达,进而介导拟除虫菊酯抗性;而外显子区域非同义替换(S162N)可能通过修饰CYP9J10基因编码的蛋白结构,增强其解毒酶活性,进而介导拟除虫菊酯抗性。以上这些P450基因的遗传变异与拟除虫菊酯杀虫剂抗性的关联性,还需进一步通过荧光素酶基因报告实验和解毒酶活性代谢实验来验证SNPs位点是否参与基因表达调控或者介导解毒酶活性。此外,本研究可能遗漏低频变异,后续需进一步扩大野外样本采集数量,以验证遗传变异的可靠性和发生频率。
-
本研究综合进化树分析、主成分分析、群体遗传结构分析结果,发现36个中华按蚊样本的聚类结果与地理分布一致,不同种群间可能存在地理隔离现象。CYP9K1、CYP6P3v1、CYP9J10的遗传变异可能介导拟除虫菊酯抗性。上述P450基因遗传变异与拟除虫菊酯抗性的关联,仍需通过荧光素酶报告基因实验和解毒酶活性代谢实验进一步验证,以明确相关SNPs能否调控基因表达或影响解毒酶活性。本研究结果为深入解析中华按蚊对拟除虫菊酯的抗性机制,特别是P450基因的表达调控与解毒酶活性提供了关键的数据基础。





 DownLoad:
DownLoad: