-
开放科学(资源服务)标识码(OSID):

-
截至2024年6月,自然界中已知的蜘蛛种类超过52 000种[1]。这些蜘蛛可以根据其生活方式大致分为3类:游猎蜘蛛、洞穴蜘蛛和结网蜘蛛。游猎蜘蛛通过四处游走或伪装捕食猎物,不结网。洞穴蜘蛛喜欢藏身于沙堆或洞穴中。结网蜘蛛的主要特征是其结网行为,首先使用丝腺合成并加工蛛丝蛋白,蛛丝蛋白被吐出后会在空气中凝结成细小的蛛丝,结网蜘蛛再用这些蛛丝结成具有高黏性的蛛网,用于捕食猎物。蜘蛛共有9种丝腺体,但并非一种蜘蛛同时拥有这9种丝腺体,如十字园蛛(Araneus diadematus)体内含有包括大囊状腺、小囊状腺、鞭毛状腺、葡萄串状腺等在内的7种丝腺体,可分泌包括蜘蛛网中呈放射状的横丝、捕食过程中缠绕食物的包裹丝以及用于制作卵囊保护后代的卵囊丝等共计7种具有不同特性和生物学功能的蛛丝[2]。
天然蛛丝大多具有卓越的机械性能,如达尔文树皮蜘蛛(Caerostris darwini)分泌的大囊状腺体丝的断裂强度可达(380±62) MPa,与高强钢相当[3]。除了优良的机械性能外,天然蛛丝良好的抗菌性、生物相容性和可降解性,使其具有成为手术缝合线、药物缓释载体等生物医用材料的潜力。然而,由于蜘蛛的独居性和肉食性,以及它们相互残杀的习性,很难进行大规模、高密度饲养,因此天然蛛丝的产量非常低,难以满足生产的需求。
随着生物技术的发展,人们对蛛丝蛋白的结构和性能有了更深入的了解,科研人员开始利用基因工程技术合成重组蜘蛛丝蛋白。1995年,Prince等[4]利用大肠杆菌(Escherichia coli)表达了分子量为14.7~41.3 kDa的蜘蛛丝蛋白,这是采用基因工程方法生产重组蜘蛛丝蛋白的最早报道。此后,研究人员陆续利用包括真菌[5-6]、植物[7-8]、哺乳动物[9-11]和家蚕细胞[12-13]或个体[14-15]等多种宿主成功表达重组蜘蛛丝蛋白。这些人工合成的蜘蛛丝蛋白不仅保留了天然蛛丝蛋白的优良特性[16],还具有更强的可塑性和可修饰性[17]。近年来,重组蜘蛛丝蛋白已经被加工成膜、微纳米球和支架等多种形态[18],在生物医学领域的应用日益广泛。本文概述了不同种类蛛丝蛋白的结构与功能,详细阐述了人工合成重组蜘蛛丝蛋白的原理和方法,并讨论其在生物医学领域的应用进展及潜在的发展前景。
HTML
-
不同种类的蜘蛛可以分泌多种不同类型的天然蛛丝,这些蛛丝又由多种不同的丝蛋白构成。2017年,Babb等[19]经全基因组测序发现,络新妇蛛(Nephila clavipes)的基因组中包含编码28种不同蛛丝蛋白的基因,其中8种是首次发现的新蛛丝蛋白。蛛丝蛋白属于高分子量蛋白(通常为250~400 kDa),其氨基酸序列根据蛛丝类型和蜘蛛种类的不同而存在差异。蛛丝蛋白通常由高度重复的序列段构成,两侧分别是非重复的氨基末端结构域(即N端结构域,NTD)和羧基末端结构域(即C端结构域,CTD)[20-21],其一级结构在不同类型的蛛丝和不同种类的蜘蛛中独一无二。进一步的序列研究表明,蛛丝蛋白的氨基酸序列和二级结构,如β-折叠、无规卷曲等,其数量、排列顺序和比例共同决定了天然蛛丝的机械性能[21]。
-
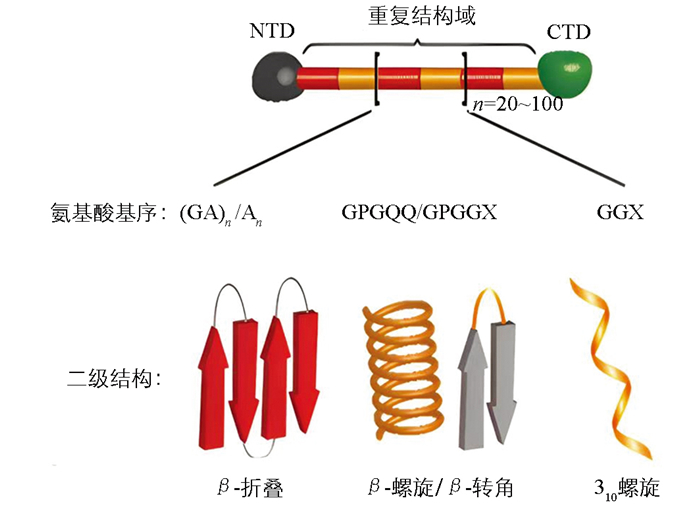
蜘蛛大囊状腺体分泌的大囊状腺体丝(Major Ampullate Gland Silk),也称为拖牵丝或牵引丝(Dragline Silk)。1990年,Xu等[22]从络新妇蛛大囊状腺体的cDNA库中成功筛选出1个长度为2.4 kb的大囊状蛛丝蛋白1(MaSp1)。随后,Hinman等[23]鉴定获得了1个富含脯氨酸的MaSp2,并推测天然蜘蛛拖牵丝主要由MaSp1和MaSp2这2种蛋白组成。进一步研究表明,这2种蛋白在不同蜘蛛拖牵丝中的比例有所不同,如金丝蜘蛛(Trichonephila clavipes)拖牵丝由81%的MaSp1和19%的MaSp2组成,而橙色金蛛(Argiope aurantia)拖牵丝中MaSp1和MaSp2的比例分别为59%和41%[24]。拖牵丝蛋白的氨基酸基序具有高度重复的特点,其重复序列主要由富含甘氨酸和丙氨酸的基序构成,如图 1。MaSp1和MaSp2由非重复的CTD和NTD以及中间的重复结构域构成,其中重复结构域通常包含约20~100个串联的重复结构单元[25]。MaSp1的重复结构域富含polyA(An)、polyGA((GA)n)和GGX(X=Q、Y、L或R)基序;MaSp2的重复结构域含有脯氨酸基序,主要为GPGQQ和GPGGX,同时也富含An基序。拖牵丝蛋白氨基酸基序组成的二级结构的数量及分布,决定了拖牵丝的力学特性。例如(GA)n/An基序参与形成的疏水性β-折叠纳米晶体结构域,起到了增强拖牵丝纤维抗拉伸强度的效果;GGX基序参与形成的亲水性310螺旋结构域,以及GPGQQ或GPGGX基序形成的β-螺旋或β-转角结构,则赋予了拖牵丝良好的弹性[25-28]。
蜘蛛拖牵丝中除了MaSp1和MaSp2这2种主要蛋白外,还含有其他含量较低的蛛丝蛋白(表 1)。1996年,Guerette等[29]利用络新妇属Spidroin1和Spidroin2的基因cDNA部分序列作为探针,从园蛛属丝腺cDNA库中筛选获得了4个阳性克隆序列,分别命名为ADF1~ADF4。研究发现,ADF3和ADF4蛋白的氨基酸序列与MaSp1和MaSp2相似,尽管这2种蛋白的脯氨酸含量较低,仅占整个氨基酸序列的13%,但无论是由ADF3/ADF4蛋白组合还是由MaSp1/MaSp2蛋白组合组成的拖牵丝,比较二者的力学性能并无显著差异。进一步研究表明,脯氨酸主要决定了蜘蛛丝的超收缩特性及湿丝的机械性能[30]。此外,研究显示超收缩性与蛋白的亲水性和疏水性有关,其中MaSp1和ADF4蛋白的疏水性较好,而MaSp2和ADF3蛋白则更为亲水[31]。
2019年,Kono等[32]利用基因组学、转录组学和蛋白质组学方法,首次从大腹园蛛(Araneus ventricosus)的蛛丝蛋白序列中鉴定获得了MaSp3,并将其命名为MaSp3A。2021年,同一研究团队又从络新妇蛛的拖牵丝中鉴定获得了MaSp3B序列[33]。2019年,Garb等[34]对达尔文树皮蜘蛛(Caerostris darwini)进行等位基因测序,获得了MaSp4和MaSp5。进一步研究发现,MaSp4中GPGQQ序列占重复序列的44%~52%,其重复序列含有31.4%~32.0%的脯氨酸,这种特殊的重复序列与蜘蛛丝优良的弹性有关,而MaSp5主要由GGX序列组成。目前,MaSp4和MaSp5这2种蛋白仅在达尔文树皮蜘蛛中发现[35]。
-
1998年,Hayashi等[47]对络新妇蛛进行了cDNA序列的克隆,首次筛选出鞭毛状腺体丝蛋白(Flagelliform Gland Spidroin,Flag)基因(表 1)。随后,Lubec团队通过蛋白质组学确定了络新妇蛛Flag的完整序列[48]。与MaSp1和MaSp2类似,Flag也拥有NTD、CTD以及由11个重复单元组成的重复结构域。这些重复单元由GPGGX、GGX(X=S、Y或A)和疏水间隔序列构成,其中GPGGX序列和GGX序列可形成310螺旋和β-螺旋,对鞭毛状腺体丝的可伸展性、韧性和强度做出了贡献[37, 49]。
2020年,Tian等[37]对大腹园蛛的Flag进行了鉴定,发现其富含甘氨酸和脯氨酸的特性,赋予了鞭毛状腺体丝极高的弹性,这是构成捕获丝核心部分的关键,但这种富含甘氨酸和脯氨酸的特性也导致Flag难以被外源表达。2022年,Jorge等[50]通过蛋白质组学分析蜘蛛鞭毛状腺体中蛋白质的表达,结果表明Flag只在鞭毛状腺体中表达,且其分子量大于250 kDa[48]。目前,由于缺乏在天然状态下Flag的真实蛋白结构数据,因此其完整结构模型尚待进一步确定[51]。
2022年,Babb等[36]在达尔文树皮蜘蛛中鉴定获得了3种新的Flag同源蛋白——Flag-A、Flag-B、Flag-E (表 1)。进一步研究发现,这3种蛋白能在雄性蜘蛛的鞭毛状腺体、毒腺、头胸部和螯肢等部位表达,而仅在雌性蜘蛛的鞭毛状腺体中表达。这与之前发现的Flag只在络新妇蛛和大腹园蛛的鞭毛状腺体中表达有所不同。此外,该研究还表明这3种Flag还与达尔文树皮蜘蛛的毒素分泌有关。
-
除了上述研究较为广泛与深入的大囊状腺体丝蛋白和鞭毛状腺体丝蛋白外,其余报道的主要蛛丝蛋白还包括小囊状腺体丝(Minor Ampullate Gland Silk)、聚集状腺体丝(Aggregate Gland Silk)、葡萄串状腺体丝(Aciniform Gland Silk)、管状腺体丝(Tubuliform Gland Silk)和梨状腺体丝(Pyriform Gland Silk)(表 1)。
小囊状腺体丝在蜘蛛的生理过程中主要起支撑和固定作用[52]。1998年,Colgin等[53]克隆获得了来自小囊状腺体丝的2种小囊状蛛丝蛋白1和2(Minor Ampullate Spidroin 1和Minor Ampullate Spidroin 2,MiSp1和MiSp2)的基因序列,其编码蛋白分子量分别为9.5 kDa和7.5 kDa。此外,小囊状腺体还会分泌ADF1,ADF1自身不具有弹性,但其能够提高蜘蛛丝的力学性能,并有助于蜘蛛丝网状结构的形成[54]。
聚集状腺体丝蛋白(Aggregate Spidroin,ASG)含有高黏性糖蛋白和高吸湿性多肽[39]。ASG1和ASG2分别在络新妇蛛和黑寡妇蛛(Latrodectus hesperus)中被鉴定获得[40]。进一步研究发现,ASG1是一种广泛存在于节肢动物的围膜基质蛋白,ASG2具有拖牵丝蛋白相似的序列和典型结构(包含NTD、重复序列和CTD),具有较强的黏附作用,是捕获丝的黏附蛋白。
葡萄串状腺体丝主要由葡萄串状蛛丝蛋白(Aciniform Spidroin,AcSp)组成,用于捕获和包裹猎物。蜘蛛葡萄串状蛛丝蛋白可以根据其含有的不同重复区域分为2种类型(AcSp1和AcSp2)[41]。AcSp由14个高度重复的同源单元组成,每个单元约含有200个氨基酸,相对于其他类型的蛛丝蛋白,AcSp中的甘氨酸和丙氨酸含量较低[27]。
管状腺体丝(Tubuliform Gland Silk)又称圆柱形腺体丝(Cylindrical Gland Silk),主要用于构建蜘蛛卵壳的外层,与内层的葡萄串状腺体丝共同构成抵御外界威胁的保护屏障[42]。Tian等[55]经克隆测序,筛选出构成管状腺体丝的主要成分蛋白TuSp1,其重复序列包含An、Sn、(SA)n、(SQ)n和GX(X=Q、N、I、L、A、V、Y、F或D)。此外,组成管状腺体丝芯纤维的主要蛋白还包括2种卵壳蛋白ECP-1和ECP-2,其分子量大小均约为100 kDa[56]。
梨状腺体丝与蜘蛛网的附着、连接及食物包覆密切相关。研究人员对蜘蛛梨状腺体丝蛋白(Pyriform Spidroin,PySp)的研究相对较少,目前仅报道鉴定获得来自黑寡妇蜘蛛的PySp1[45]和来自络新妇蜘的PySp2[46]。
1.1. 蜘蛛大囊状腺体丝蛋白的结构及功能
1.2. 鞭毛状腺体丝蛋白的结构及功能
1.3. 其他主要蛛丝蛋白的结构及功能
-
目前,人工合成重组蜘蛛丝蛋白的方法主要包括化学模拟合成(Chemical Mimic Synthesis)和基因工程(Genetic Engineering)两种。化学模拟合成法是以动物丝氨基酸序列结构相似的片段为目标,通过诱导变形的方法合成重组蜘蛛丝蛋白[57]。基因工程法则是通过转基因技术将蛛丝蛋白基因导入大肠杆菌、酵母等微生物,或烟草、马铃薯、家蚕、仓鼠细胞等植物和动物个体或细胞等宿主体内,以实现重组蜘蛛丝蛋白的高量表达[16](表 2)。
-
蛛丝蛋白的化学模拟合成主要采用N-羧基环内酸酐(N-carboxyanhydride,NCA)合成法和液相多肽合成法。NCA合成法在20世纪被广泛用于制备高分子量聚氨基酸,其是利用α-氨基酸环化生成NCA,再通过亲核试剂或碱催化的开环聚合方法。曾有研究利用丙氨酸-NCA开环聚合模拟蛛丝蛋白,虽然此法操作简单、反应快、产物分子量大,但所得到的重组蜘蛛丝蛋白结构与天然蛛丝蛋白相差较大[58]。Fukushima等[59]利用五氯本酯活化酯的液相多肽合成法,模拟合成了分子量在23~48 kDa的重组蜘蛛丝蛋白,其含有多个连续重复的GGX(X=Ala、Tyr、Gln或Leu)氨基酸序列,通过甲酸溶解检测其二级结构,发现这些重复序列蛋白形成了β-折叠结构。然而,由于液相多肽合成法步骤繁琐,聚合物分子量较小,因此该方法目前主要用于研究模拟蛛丝蛋白与天然蛛丝蛋白结构及性能之间的关系,并未广泛应用于实际开发。
近年来,研究人员还致力于利用化学合成法创制具有优异力学性能的人造蜘蛛丝。2019年,Dou等[60]基于天然蛛丝的壳-芯结构,使用乙烯基三乙氧基硅烷、丙烯酸及去离子水合成了聚丙烯酸水凝胶,再通过水蒸发自组装方式,利用该水凝胶进一步获得了机械性能可与天然蛛丝相媲美的人造蜘蛛丝纤维。上述方法为人造蜘蛛丝纤维的合成开辟了一条新途径,但通过自组装获得具有更为优异力学性能的人造蜘蛛丝仍具挑战。2024年,He等[61]通过施加外部应力来诱导自组装过程,优化了蛋白分子链的柔韧性,成功利用超细纳米原纤维制备更坚固的人造蜘蛛丝,其具有1.83 GPa的断裂强度以及高达238 MJ/m3的韧性。
-
1995年,Prince等[4]首次使用大肠杆菌表达了分子量在14.7~41.3 kDa的蜘蛛丝蛋白。1996年,Lewis等[62]利用大肠杆菌表达了分子量超过100 kDa的蜘蛛丝蛋白,其质量分数在分泌总蛋白中达到10 mg/g。1997年,Fahnestock等[63]分别以大肠杆菌和毕赤酵母(Pichia pastoris)作为宿主合成了重组蜘蛛丝蛋白DP-1B,结果发现大肠杆菌只能有效合成并分泌小于1 000个氨基酸的蜘蛛丝蛋白,而毕赤酵母能够合成并分泌含有3 000个及以上氨基酸的蜘蛛丝蛋白,但其表达效率随氨基酸数量增加而降低。2010年,Xia等[64]利用大肠杆菌表达了分子量为284.9 kDa的重组MaSp1,通过将重组MaSp1溶解在常用的纺丝溶剂六氟异丙醇中,随后以溶解后浓度为20%的重组MaSp1溶液为原料,经电纺织成重组蜘蛛丝纤维,检测其力学性能可达到蜘蛛天然拖牵丝相当的水平。2016年,Jansson等[6]利用毕赤酵母生产了重组蜘蛛丝融合蛋白Z-4RepCT,通过蛋白质自组装可变成丝纤维。值得注意的是,由于蛛丝蛋白富含甘氨酸和丙氨酸且具有复杂的二级结构,利用大肠杆菌和酵母等微生物表达重组蜘蛛丝蛋白时,常会出现遗传不稳定的现象,导致重组蜘蛛丝蛋白表达效率降低[65]。
近年来,研究人员还尝试利用其他微生物合成重组蜘蛛丝蛋白。2020年,Foong等[65]利用海洋光合紫色细菌(Rhodovulum sulfidophilum)成功表达了源自络新妇蛛的重组MaSp1,与大肠杆菌相似,海洋光合紫色细菌也会面临因遗传不稳定而导致重组蜘蛛丝蛋白表达效率低的问题。2022年,Jin等[66]利用谷氨酸棒状杆菌(Corynebacterium glutamicum)成功合成并分泌了2种源自络新妇蛛的重组MaSp1,其分子量大小分别为43 kDa和168 kDa,遗憾的是通过微流体纺纱装置获得蛛丝纤维的力学性能未能达到天然蛛丝的水平。
-
除了在微生物中表达重组蜘蛛丝蛋白外,研究人员还曾利用多种植物作为宿主表达重组蜘蛛丝蛋白。2001年,Scheller等[7]合成了源自络新妇蛛包含有420~3 600个核苷酸的MaSp1的cDNA,该基因序列与天然蛛丝蛋白基因序列具有高度相似性,随后利用烟草和马铃薯作为宿主表达该融合基因,但重组蜘蛛丝蛋白仅占所有蛋白含量的2%[7]。2013年,Hauptmann等[67]利用基于内含肽介导的翻译后蛋白融合技术(Intein-Based Posttranslational Protein Fusion Technology)在烟草叶片细胞的内质网中添加了络新妇蛛的Flag基因序列,并成功获得了分子量超过250 kDa的Flag。2016年,Peng等[8]也利用该技术在烟草中成功表达了分子量为73~136 kDa的重组MaSp1和MaSp2。2008年,Brooks等[68]在拟南芥的叶片和种子及大豆胚中成功表达了源自络新妇蛛的重组MaSp2。与微生物相比,植物表达重组蜘蛛丝蛋白的优势在于其可以利用初始原材料自主合成氨基酸,且重组率更高、遗传稳定性更强,但还存在以下问题:通过植物生产蜘蛛丝蛋白需要全面且昂贵的临床研究,植物和植物细胞的生产效率低,纯化重组蜘蛛丝蛋白成本高等,因此还需要进一步完善技术以便实际应用[67, 69]。
2002年,加拿大Nexia生物技术有限公司与美国陆军Natick研究中心共同完成了利用哺乳动物来表达重组蜘蛛丝蛋白的研究[9]。他们首先合成了源自络新妇蛛牵引丝的MaSp1和ADF3的cDNA,将其分别在牛乳腺上皮泡状细胞和仓鼠肾细胞中进行表达,并将纯化后的蛛丝蛋白纺丝得到了世界上首例通过哺乳动物合成的高弹性、高强度的人造蜘蛛丝纤维,称为“生物钢”。2007年,Xu等[70]将人工合成的络新妇蛛的MaSp1和MaSp2蛋白基因插入羊的β酪蛋白启动子序列下游,注射进小鼠受精卵的细胞核中,最终在小鼠的乳汁中成功表达了重组蜘蛛丝蛋白。此后,也有研究尝试利用山羊、猪等动物表达蛛丝蛋白[16]。与微生物或植物的表达系统不同,重组蜘蛛丝蛋白在动物体内的表达并不是独立存在,而是与动物本身的酪蛋白或乳清蛋白均匀混合,因此如何进行有效的分离和回收是动物表达重组蜘蛛丝蛋白的关键。2021年,Li等[71]成功制备了自身基因组中整合有重组蜘蛛丝蛋白基因的转基因羊胚胎。与通过乳腺分泌重组蛋白不同,该研究旨在利用转基因羊丰富的毛发和毛囊生产重组蜘蛛丝蛋白,然而,尽管进行了胚胎移植并观察到妊娠,但并未产生后代,因此,通过毛发生产重组蜘蛛丝蛋白的可行性尚未得到验证[71]。
-
相较于同类相食的蜘蛛,家蚕可以进行大规模的人工饲养,能合成大量蛋白质并纺丝结茧。家蚕的丝腺与蜘蛛的丝腺形态十分相似[72],且它们的理化环境表现出极高的相似性[14]。蜘蛛与家蚕的丝蛋白均由富含甘氨酸和丙氨酸的串联重复序列组成,两者的基因序列和结构也高度相似[72-73]。经过人类数千年的驯化,家蚕已能高效合成丝蛋白,因此借助于转基因或基因组编辑等手段创制遗传改良家蚕,进而实现在遗传改良茧丝中高效合成重组蜘蛛丝蛋白的方法具有其他物种不可比拟的优势。
1999年,Altmann等[13]首次利用家蚕核型多角体病毒杆状表达系统在家蚕细胞中成功表达了37 kDa的重组蜘蛛鞭毛状蛋白,随后Zhang等[15]也利用此系统在家蚕细胞系(BmN)和幼虫中成功表达了70 kDa的MaSp1-EGFP荧光融合蛋白。研究报告称,在幼虫中产生的融合蛋白产量为6 mg,占BmN细胞总蛋白的5%。此外,该研究指出,重组蜘蛛丝蛋白的低溶解性是产量提升的主要障碍[15]。近年来,研究人员开始通过利用piggyBac转座载体建立更为可靠的转基因家蚕来表达重组蜘蛛丝蛋白[74-78]。Wen等[75]获得了基因组中含有家蚕丝胶蛋白1(Sericin 1)基因启动子调控MaSp1表达框的转基因家蚕,并成功在转基因茧丝中检测到了MaSp1的表达。然而,由重组蜘蛛丝蛋白纺成的丝蛋白纤维力学性能不如天然蜘蛛拖牵丝,这可能是由于MaSp1在丝蛋白中占比较低[75]。2014年,Kuwana等[76]构建了拥有大腹园蛛拖牵丝蛋白基因和家蚕丝素重链基因融合片段的转基因家蚕,并获得了含有融合蛋白的转基因蚕丝。经检测,转基因蚕丝的韧性与非转基因蚕丝相比提高了53%,随后这种转基因纤维被编织成背心和围巾等衣物。2018年,You等[77]利用piggyBac转座载体成功建立了能表达‘黑寡妇’体内大小为160 kDa的MaSp2转基因纯合性家蚕和杂合性家蚕,通过对两种家蚕丝纤维的力学性能比较,纯合性家蚕的丝纤维在最大应力、杨氏模量、韧性等方面均优于杂合性家蚕。2021年,Tang等[78]也利用该转座子系统,成功获得了2种转基因家蚕,能分别在茧丝中表达金丝蜘蛛ASG1和金银蛛(Argiope argentata)的PySp1。该研究还表明,这2种蛋白均能提高转基因蚕丝的力学性能,其中含ASG1的茧丝韧性和断裂拉升强度与天然茧丝相比分别提高了21.0%和34.2%,含PySp1的茧丝韧性和断裂拉升强度分别为91.5%和36.9%。
近年来,研究人员开始探索利用基于转录激活因子样效应核酸酶(TALEN)或CRISPR/Cas9的基因组编辑工具,将外源重组蜘蛛丝蛋白基因与内源丝蛋白基因进行原位替换,期望进一步提高重组蜘蛛丝蛋白在遗传改良家蚕茧丝中的含量。2018年,Xu等[79]利用TALEN介导的外源基因替换技术,用络新妇蛛的MaSp1蛋白基因取代了家蚕丝素重链基因,获得的遗传改良茧丝中含有35.2%的重组MaSp1。次年,Zhang等[80]利用CRISPR/Cas9技术成功将络新妇蛛MaSp1和MiSp1蛋白基因插入到丝素蛋白重链或轻链中,得到的蛋白纤维拉伸强度与天然蛛丝相当,且能保持遗传稳定性。2023年,Mi等[14]也利用CRISPR/Cas9技术成功地在转基因家蚕中合成了达尔文树皮蜘蛛完整的聚酰胺蜘蛛丝纤维,该纤维的拉伸强度达到1 299 MPa,韧性达到319 MJ/m3,该研究对具有高强度和超韧性的商用纤维生产具有重要参考意义。2024年,Yu等[81]利用3种不同转化策略:转座子介导的转基因重链表达系统(BmFibH-T)、TALEN介导的丝素蛋白轻链融合系统(BmFibL-F)和丝素蛋白重链替换系统(BmFibH-R),获得了丝腺能特异表达源自络新妇蛛的MaSp2并将其分泌至茧丝的转基因家蚕中。结果显示,与天然茧丝相比,利用BmFibL-F、BmFibH-R和BmFibH-T获得的转基因茧丝的断裂拉伸强度分别提高了34.86%、73.87%和86.18%,其中由BmFibH-R构建的转基因蚕丝中的重组外源MaSp2的含量最高(占茧丝总蛋白的51.02%),表明利用BmFibH-R建立转基因家蚕可能是大量生产重组蜘蛛丝蛋白的有效手段[81]。
2.1. 化学模拟合成重组蜘蛛丝蛋白
2.2. 微生物合成重组蜘蛛丝蛋白
2.3. 植物和哺乳动物合成重组蜘蛛丝蛋白
2.4. 遗传改良家蚕“纺”蜘蛛丝
-
数千年来,人类利用蜘蛛丝作为各种用途的天然材料。在古代,希腊人和罗马人使用以天然蛛丝蛋白为原料的膏药作为敷料用于战伤止血,这些膏药还被用作止疼药和退烧药[90]。然而,由于从蜘蛛中直接提取蛛丝极度耗时,再加上蜘蛛的自相残杀特性导致大规模饲养蜘蛛难度极大,因此从自然界中收集蛛丝极不容易,无法提供大量的材料。
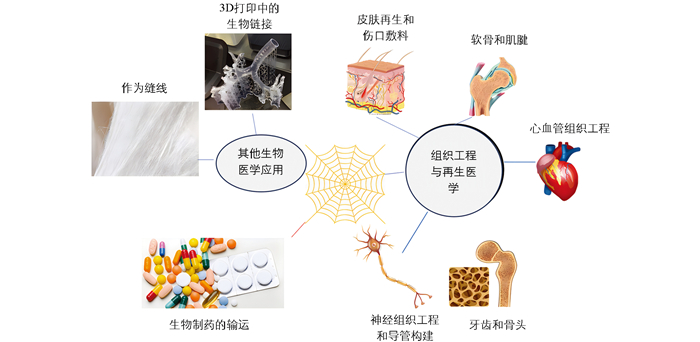
近年来,随着生物技术的发展,重组蜘蛛丝蛋白的产量增加且质量更为一致,一些重组生产的蜘蛛丝变种甚至已经商业化[91]。研究人员通过大量的细胞体外试验发现,溶液状态的重组蜘蛛丝蛋白大多无细胞毒性,且重组蜘蛛丝蛋白生物材料具有良好的生物相容性、生物降解性、低过敏性和可修饰性。此外,重组蜘蛛丝蛋白还具有易加工性,可进一步开发为薄膜(Film)、微纳米球(Micro-nanosphere)、支架(Scaffold)、纤维(Fiber)、凝胶(Hydrogel)、胶囊(Capsule)和网(Net)等结构(图 2),因此其在生物医学领域备受青睐[92-96]。
-
为了制备具有多种不同形态结构的重组蜘蛛丝蛋白材料,研究人员首先利用不同的缓冲液溶解重组蜘蛛丝蛋白,然后将溶液置于模具中自然风干,从而获得具有所需形态结构的产物[97],利用这种方式制备的重组蜘蛛丝蛋白膜具有优异的机械特性和选择渗透性。与天然蛛丝制成的蛋白膜相比,重组蜘蛛丝蛋白膜更薄,更适合用作医学材料[98]。这种重组蜘蛛丝蛋白膜可用于固定和包裹酶、抗体、抗原以及药物等,进而制成抗菌材料[99]以及伤口敷料[100],并能在组织再生领域[101]和药物递送领域[102]发挥作用。
将蛛丝蛋白与抗菌多肽结合,可以得到新型的抗菌材料。2011年,Gomes等[103]将人中性粒细胞防御素-2(HNP-2)以及人中性粒细胞防御素-4(HNP-4)与络新妇蛛的重组MaSp1结合得到复合蛋白膜。同年,Currie等[99]将短银结合肽对应的核苷酸整合到络新妇蛛的基因组中,再用其表达的重组蜘蛛丝蛋白制成的膜分别去处理大肠杆菌和金黄色葡萄球菌,结果表明大肠杆菌和金黄色葡萄球菌的生长均受到抑制。此外,重组蜘蛛丝蛋白膜作为植入物涂层,可以有效减少细菌感染,并能通过改善细胞附着和生长状态来优化植入物的性能以及与人体的整合[104]。
在古代,天然蛛丝被用来覆盖伤口并帮助伤口愈合,如今对蛛丝敷料的研究也表明其具有治愈烧伤伤口的潜力[91],因此重组蜘蛛丝蛋白膜还可用于伤口敷料。Zhao等[100]利用pNSR16/PVA静电纺丝膜为敷料治疗大鼠皮肤烧伤,其结果表明重组蜘蛛丝蛋白膜能促进大鼠受损皮肤的愈合。Chouhan等[105]将涂有和未涂有重组蜘蛛丝蛋白的薄膜,分别用于兔子糖尿病伤口,结果表明涂有重组蜘蛛丝蛋白的薄膜能加速兔子伤口的愈合。
重组蜘蛛丝蛋白膜也被广泛应用于促进组织再生的研究中。有报道表明,重组蜘蛛丝蛋白膜的光滑面能有效将骨缺损部位与非成骨细胞隔离,而其粗糙面则有助于促进成骨细胞的黏附和增殖[106]。重组蜘蛛丝蛋白膜对人体血浆蛋白具有渗透性,并促进人角质细胞的黏附和增殖,在膜两侧接种的人角质细胞在3 d内可形成汇合的单层[97]。此外,被重组蜘蛛丝蛋白修饰过的膜蛋白也能促进组织再生。Widhe等[107]对成纤维细胞、角质形成细胞、内皮细胞和神经胶质细胞的研究显示,经过RGD修饰的重组蜘蛛丝蛋白4RepCT在这些细胞中的黏附性更加显著,更能促进组织再生。随后,Schiller等 [108]研究表明,eADF4(C16)-RGD膜与eADF4(C16)膜相比,含有重组蜘蛛丝蛋白的前者更能促进成纤维细胞的黏附和分裂。
经生物工程改造的重组蜘蛛丝蛋白薄膜,最近也被开发成药物输送系统[109]。有研究表明,由于阳离子药物优先与多阴离子的eADF4(C16)相互作用,可通过调整eADF4(C16)在薄膜中的比例,来调节重组蜘蛛丝蛋白膜的药物加载量[110]。该研究还表明这些薄膜可以在几周内有效地调控药物释放,并在弹性蛋白酶和胰蛋白酶存在时加速药物释放,类似于家蚕丝素薄膜的药物释放特性,特别适合递送如氟哌啶醇(Haloperidol)之类的阳离子抗精神病药物。然而,利用重组蜘蛛丝蛋白薄膜构成的药物递送系统并不利于药物的包裹[91],因此仍需进一步研究。
-
重组蜘蛛丝蛋白微纳米球的制备方法更具多样性和复杂性,包括乳液法、自组装法、微制备法、离子凝聚法、相分离法等[111]。封装药物并进行药物递送,是重组蜘蛛丝蛋白微纳米球最为广泛的应用方式[112]。将重组蜘蛛丝蛋白溶液与药物混合,加入甲醛、六氟异丙醇等,蛋白与药物会发生自组装,形成微米或纳米颗粒,经水或乙醇处理后性质趋于稳定,可用于后续研究[110, 113]。Jastrzebska等[114]将重组蜘蛛丝蛋白微纳米球作为抗癌药物的载体,结果表明这种微纳米球具有肿瘤组织的靶向性,可延长药物的释放时间,从而增强了药物对肿瘤细胞的作用效果并降低了对正常细胞的损害。该研究还表明,重组蜘蛛丝蛋白微纳米球由于其优良的弹性和机械性能,可以确保封装药物的稳定性,而且这种材料可以调控封装药物在病灶部位的精准释放剂量。Gu等[115]对重组蜘蛛丝蛋白微纳米球递送疏水和亲水性药物进行了研究,结果发现重组蜘蛛丝蛋白微纳米球能够稳定包裹疏水性药物并能将其准确递送至靶细胞或靶器官,进而发挥药效。此外,由于制备的重组蜘蛛丝蛋白微纳米球带有负电荷,其可与带正电荷的亲水性药物通过静电作用相结合形成稳定的聚合物,进而将其递送至肿瘤组织发挥药效。
重组蜘蛛丝蛋白微纳米球本身还具有促进细胞生长增殖的功效。2022年,Deptuch等[116]将重组蜘蛛丝蛋白微纳米球用于刺激小鼠脾细胞,其结果表明被刺激过的小鼠脾细胞增长速率更快,说明重组蜘蛛丝蛋白微纳米球有促进小鼠细胞生长的功能。2023年,Ornithopoulou等[117]将重组蜘蛛丝蛋白微纳米球接种于人骨髓间充质干细胞,微纳米球能黏附在细胞表面,随后细胞随着微纳米球逐渐融合在一起,其结果表明重组蜘蛛丝蛋白微纳米球能促进人细胞的生长。
-
生物支架既要能起到支撑作用还需要为细胞的黏附、分化和繁殖提供场所。最常用制备支架的方法是冻融法:将重组蜘蛛丝蛋白溶液浇铸在适当模具中,低温处理一定时间,室温解冻后并用水冲洗即得到支架材料[118]。2024年,Branković等[119]利用NIH-3T3细胞对重组蜘蛛丝蛋白支架进行了体外测试,结果证实NIH-3T3细胞可以在重组蜘蛛丝蛋白支架上轻松附着并不阻碍其生长,表明重组蜘蛛丝蛋白支架具有良好的细胞相容性。此外,利用重组蜘蛛丝蛋白制得的3D支架可以定制,其可以通过控制营养物质和氧气的运输,实现对细胞迁移、增殖和分化的控制[106],能在生物体内进一步开发利用。
重组蜘蛛丝蛋白支架具有再生生物医学应用的良好特性,可用于组织修复以及伤口敷料等[120]。利用重组蜘蛛丝蛋白制得的3D支架力学性能良好,足以支撑起骨组织,可以用作修复骨组织的材料[121]。重组蜘蛛丝蛋白支架还能构成人体血管,可用于血管移植[122]或血管再生[123]。重组蜘蛛丝蛋白还能被用于构成伤口愈合支架,促进皮肤再生[124]。在Chouhan等[125]和Liu等[126]的研究中,将来自蚕的丝素蛋白加工成纳米支架,并用重组蜘蛛丝蛋白(FN-4RepCT)修饰,利用该支架覆盖大鼠模型中的三级烧伤伤口,经过14 d的治疗后,与利用市售的敷料贴片处理的伤口以及未经治疗的伤口相比,这种功能性支架显示出更好的促伤口愈合的功效。
-
重组蜘蛛丝蛋白形成的纤维在医学领域最主要的用途是用作手术缝合线[90]。2013年,Hennecke等[127]将重组蜘蛛丝蛋白纤维制备的手术缝合线与天然蛛丝制得的传统手术缝合线进行比较,结果证明2种手术缝合线的抗拉强度基本相同,重组蜘蛛丝蛋白制备的手术缝合线在1 000个疲劳周期内的强度并不会显著降低。此外,重组蜘蛛丝蛋白形成的亚微米和纳米纤维也可用于再生医学,如膜组织[128]、神经导管[129]、皮肤移植物和伤口敷料等[130]。
重组蜘蛛丝蛋白水凝胶的形成过程是在再生丝蛋白溶液中加入水溶性有机物或无机物形成纳米纤维,然后进一步浇铸成型。水凝胶不同于支架等材料,它不存在孔径间隙,其结构中充满了大量水分。重组蜘蛛丝蛋白形成水凝胶能够封存细胞,随后可以在其3D结构中处理细胞,进行3D生物打印[96, 104]。此外,将重组蜘蛛丝蛋白与其他生物活性化合物如海藻酸盐结合的水凝胶作为伤口敷料,已经在促伤口愈合研究方面显示出良好的活性[119]。
重组蜘蛛丝蛋白在甲苯中可以快速形成微米级的胶囊,可用作药物的递送载体,药物通过蛛丝蛋白的水解而释放,适应于靶向药物运输[131]。重组蜘蛛丝蛋白微针已作为特殊材料被用于胰岛素的药物递送中[132]。2019年,Pawar等[51]评估了基于重组蜘蛛丝蛋白的无纺布网应用于神经修复领域的效果,结果表明该无纺布网的表面结构能为神经再生提供理想的附着位点,且没有细胞毒性。
3.1. 重组蜘蛛丝蛋白膜
3.2. 重组蜘蛛丝蛋白微纳米球
3.3. 重组蜘蛛丝蛋白支架
3.4. 其他形态的重组蜘蛛丝蛋白
-
近年来,研究人员已经不限于探究利用不同的合成策略来生产重组蜘蛛丝蛋白,还在努力改良和优化重组蜘蛛丝蛋白序列的方法以及与其他功能性分子联合使用的策略,目前已经有一系列由不同宿主合成的重组蜘蛛丝蛋白被研究,其表达得率高,能满足探索下游材料潜在应用开发的需求。研究人员将注意力转向在重组蜘蛛丝蛋白氨基酸序列中插入一些特殊的结构序列,从而对蛛丝蛋白分子的构象进行人工调节并赋予其新性能,以期应用于生物医学领域。具有特异性功能的蛋白或核酸序列,如可控细胞穿透与细胞膜失活肽(Cell Penetrating and Cell Membrane Destabilizing Peptides,CPPs)、核酸适体(Aptamer)等,也有望应用于制成蛛丝蛋白新型材料[133]。此外,重组蜘蛛丝蛋白与离子化合物、功能性肽等结合后也具有特殊功能,如增强细胞黏附性、促进细胞生长增殖、特异性识别靶位点和抗菌等[93, 134]。
重组蜘蛛丝蛋白还被探索用于开发各种新型绿色能源转换设备及潜力巨大的生物医用材料,如植入式器官监护仪[16]。研究显示,通过在重组蜘蛛丝蛋白中添加特定的信号传导和黏附基序,可以促进干细胞的分化,或者通过调控其生物降解性来构建人造骨等永久性植入物,以及生产安全的唇部用药和烧伤愈合产品[95]。由于重组蜘蛛丝蛋白具有优良的生物相容性,它可用于开发药物递送研究和毒素测试中的器官芯片[135],或制造人工假体、韧带和皮肤等人造材料[115]。结合其低免疫反应,这些材料在细胞培养和药物递送应用领域的研究进展顺利[136]。
重组蜘蛛丝蛋白的这些优良特性和潜在的应用价值都需要在未来进行广泛研究,以提供合适且安全的蜘蛛丝基生物材料,用于药物递送和再生医学等领域。然而,重组蜘蛛丝蛋白的利用仍面临制备工艺优化和材料应用安全性能等方面的挑战,需要更多研究来保持重组蜘蛛丝蛋白材料的稳定性和安全性[95]。
-
现有研究表明,蛛丝蛋白种类丰富且具有优良的力学性能,为其在医学领域的多功能应用奠定了基础,但由于天然蛛丝蛋白无法量产,通过人工手段合成重组蜘蛛丝蛋白成为推动其在医学领域实现大规模应用研究的关键环节。重组蜘蛛丝蛋白可以通过化学合成和基因工程技术获得,并通过各种加工技术开发成适用于不同医学领域的材料,在伤口敷料、组织再生、药物递送等方面发挥着重要作用,且其应用范围还在不断拓展,但重组蜘蛛丝蛋白的研究仍面临工艺优化和材料应用安全性等问题,还需进一步研究。





 DownLoad:
DownLoad: